The Influence of Gene Polymorphisms on Tobacco and Alcohol-Induced Oral Cancer RisK
Posted by Acubiz | BlogAuthor(s)
Otávio A. Curioni, Marcos B. de Carvalho, Rogério A. Dedivitis, Abrão Rapoport, Gilka J. F. Gattas
ABSTRACT
Aims: This study examined whether genetic polymorphisms of tobacco and alcohol-related metabolic genes such as GSTM1, GSTT1, GSTP1, CYP1A1, CYP2E1 and DNA repair genes (XRCC1 194Trp, XRCC1 399Gln, andXRCC3 Met) contribute to the risk of developing OSCC.
Methods: Patients eligible for inclusion were over 18 years, had pathologically confirmed OSCC and were followed prospectively for at least two years or until death, from December 2000 to December 2004. Ninety-two OSCC patients were included along with 244 subjects from the same hospital, evaluated in the same period as patients without cancer, as the control group.
Results: GSTM1 null and XRCC1-194Trp alone increased the risk of OSCC (OR, 2.15; 95% CI, 1.2 – 3.6 and OR, 2.02; 95% CI, 1.01 – 4.03, respectively). The joint effect of GSTM1 null with CYP1A1 or CYP2E1 polymorphism increased the risk two to threefold. Similar results were observed when XRCC1-194Trp was combined with GSTM1 null or the CYP2E1 polymorphism. By contrast, XRCC1- 399Gln was associated with protection against OSCC. Gene-gene and gene-environmental interactions were mainly detected for CYP1A1 andGSTP1 associated with more than 20 p/y of tobacco and XRCC1-194Trp when more than 30 g/L/d of alcohol was consumed (OR, 8.8; 95% CI; 1.3 – 45.7).
Conclusions: The drug metabolizing and DNA repair enzyme polymorphisms may be informative for clinicians in the preventive management of patients at risk, particularly those with strong smoking and drinking habits.
Background: Scoliosis is a common condition in adult patients, and often causes chronic back pain compared to non-scoliosis. It has also been identified that bone mineral density is very often compromised in patients with scoliosis, even occurring in adolescence. At this time, no study has looked at how bone mineral density and scoliosis incidence or severity may be connected with specific regard to treatment. This study presents data on the outcomes of a scoliosis-specific exercise therapy and its ability to correct scoliotic curvatures in adult patients alone and in combination with bone mineral density supplementation. Methods: The charts of 14 total patients were retrospectively selected based upon specific inclusion criteria. Outcome assessments included the radiographic Cobb angle of the primary curvature, as well as laboratory measures of urinary deoxypyridinoline cross links. These results were compared against 12 patients who did not take the bone density supplement during or after their exercise-based treatment. Results: Patients taking the bone density supplement achieved the same level of Cobb angle reduction as compared to the control group. However, they additionally achieved a significant reduction in urinary deoxypyridinoline cross links as compared to the control group at 6 months. Conclusion: Patients taking a multi-ingredient bone density supplement daily for 6 months after completing a scoliosis-specific exercise program reported statistically significant improvements in urinary deoxypyridinoline cross links as compared to controls. It is unknown if or how bone density loss may contribute to the onset or progression of scoliosis. Long-term follow-up of these patients will be ongoing to assess bone mineral density status and Cobb angle changes longitudinally.
A Candid Assessment of the Link between Oral Candida Containing Biofilms and Oral Cancer
Posted by Acubiz | BlogAuthor(s)
J. Kang, Y. He, D. Hetzl, H. Q. Jiang, M. K. Jun, M. S. Jun, M. Khng, N. Cirillo, M. J. McCullough*
Affiliation(s)
Melbourne Dental School, The University of Melbourne, Parkville, Australia.
ABSTRACT
An association between Candida and oral cancer has long been discussed in the literature and in particular Candida albicans has been proposed to play a significant role in the development of oral cancer. Although this organism is a common member of the oral mucosa and can cause a variety of oral mucosal lesions, the exact mechanisms by which C. albicans potentially causes the development of malignant disease still require much research. We have undertaken an extensive literature search to understand pathogenicity of C. albicans, including its virulence factors, its interactions with the host immune, how a dysregulation of the immune response can result in malignancy, and how a potential C. albicans specific cytokine response may be involved in oral carcinogenesis. We present here a candid assessment of the role of Candida in oral carcinogenesis and a hypothetical model about how this may occur. Finally, we present potential future research which is necessary to elucidate.
High-fat diet linked to intestinal stem cell changes, increased risk for cancer
Posted by Acubiz | BlogOver the past decade, studies have found that obesity and eating a high-fat, high-calorie diet are significant risk factors for many types of cancer. Now, a new study from Whitehead Institute and MIT’s Koch Institute for Integrative Cancer Research reveals how a high-fat diet makes the cells of the intestinal lining more likely to become cancerous.
The study of mice suggests that a high-fat diet drives a population boom of intestinal stem cells and also generates a pool of other cells that behave like stem cells — that is, they can reproduce themselves indefinitely and differentiate into other cell types. These stem cells and “stem-like” cells are more likely to give rise to intestinal tumors, says Omer Yilmaz, an MIT assistant professor of biology and co-leader of the research team.
“Not only does the high-fat diet change the biology of stem cells, it also changes the biology of non-stem-cell populations, which collectively leads to an increase in tumor formation,” says Yilmaz, who is a Koch Institute member and a gastrointestinal pathologist at Massachusetts General Hospital.
“Under a high-fat diet, these non-stem cells acquire the properties of stem cells so that when they are transformed they become tumorigenic,” says Whitehead Member David Sabatini, who is also an MIT professor of biology and an investigator of the Howard Hughes Medical Institute.
Sabatini and Yilmaz, who previously collaborated on research into the effects of caloric restriction on stemness in the intestine, are the senior authors of the study, which appears in Nature on March 2.
People who are obese have a greater risk of developing colorectal cancer, according to previous studies. Sabatini and Yilmaz, whose labs study the relationship between diet and cancer, set out to uncover the cellular mechanisms underpinning the enhanced risk of colon cancer.
“We wanted to understand how a long-term high-fat diet influences the biology of stem cells, and how such diet-induced changes that occur in stem cells impact tumor initiation in the intestine,” Yilmaz says.
Recent studies have shown that intestinal stem cells, which last a lifetime, are the cells most likely to accumulate the mutations that give rise to colon cancer. These stem cells live in the lining of the intestine, known as the epithelium, and generate all of the different cell types that make up the epithelium.
To investigate a possible link between these stem cells and obesity-linked cancer, researchers fed healthy mice a diet made up of 60 percent fat for nine to 12 months. This diet, according to the scientists, is much higher in fat than the typical American diet, which is usually about 20 to 40 percent fat.
During this period, the mice on the high-fat diet gained 30 to 50 percent more body mass than mice fed a normal diet, and they developed more intestinal tumors than mice on a normal diet.
These mice also showed some distinctive changes in their intestinal stem cells, the researchers discovered. First, they found that the mice on a high-fat diet had many more intestinal stem cells than mice on a normal diet. These stem cells were also able to operate without input from neighboring cells.
Normally, intestinal stem cells are surrounded by support or “niche” cells, which regulate stem cell activity and tell them when to generate stem cells or differentiated cells. However, the stem cells from mice on a high-fat diet were more able to function on their own. When these stem cells were removed from the mice and grown in a culture dish without their niche cells, they gave rise to “mini-intestines” much more readily than intestinal stem cells from mice on a normal diet.
“You have more stem cells and they’re able to operate independent of inputs coming from their microenvironment,” Yilmaz says.
The researchers also found that another population known as progenitor cells — differentiated daughter cells of stem cells — started to behave like stem cells: They began to live much longer than their usual lifespan of a few days, and they could also generate mini-intestines when grown outside of the body.
“This is really important because it’s known that stem cells are often the cells in the intestine that acquire the mutations that go on to give rise to tumors,” Yilmaz says. “Not only do you have more of the traditional stem cells (on a high-fat diet), but now you have non-stem-cell populations that have the ability to acquire mutations that give rise to tumors.”
The researchers also identified a nutrient-sensing pathway that is hyper-activated by the high-fat diet. The fatty acid sensor, known as PPAR-delta, responds to high levels of fat by turning on a metabolic process that enables cells to burn fat as an energy source instead of their usual carbohydrates and sugars.
“Indeed, small-molecule agonists of PPAR-delta mimic the effects of a high-fat diet in animals fed a normal diet,” Sabatini says.
In addition to activating this metabolic program, PPAR-delta also appears to turn on a set of genes that are important for stem cell identity, Yilmaz says. His lab is now further investigating how this happens in hopes of identifying possible cancer drug targets for tumors that arise in obesity.
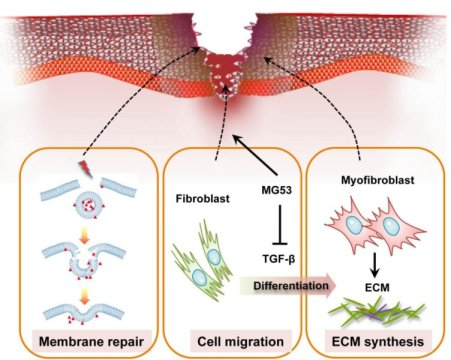
Gene identified that helps wound healing
Posted by Acubiz | BlogThis image shows modulation of wound healing and scar formation by MG53-mediated cell membrane repair and TGF-? signaling regulation.
Credit: Li.et.al., 2015
Researchers at Ohio State University have pinpointed a human gene product that helps to regulate wound healing and may control scarring in people recovering from severe injuries and damage to certain internal organs.
The protein, MG53, travels throughout the bloodstream and helps the body fix injuries to the skin, heart, lungs, kidneys and other organs without causing scars. It’s a discovery that could help heal open wounds, decrease recovery time after surgery and reduce the spread of infections.
“A massive scar on your skin may look bad, but imagine you have a heart attack and get a scar on your heart–that could be lethal,” says Jianjie Ma, a physiologist at Ohio State and co-author of the presentation.
All animals carry this gene, he said, and it’s almost identical no matter which species. MG53 fixes the cell and tissue damage that occurs during everyday living. Even simple actions, like walking or typing, will cause injuries to the body. Usually this isn’t a problem because MG53 can make repairs before there’s any serious harm.
Ma and his team genetically engineered mice without the gene that makes MG53 to see what would happen without its healing capabilities. The experiments showed that the mice lacking MG53 had difficulty recovering from injury, because of their compromised repair capacity; their heart would not function well under stress conditions.
MG53 works in tandem with another protein called TGF Beta, a type of “cytokine” protein that also heals wounds, but the healing process happens so quickly that it causes scars. If you have more TGF Beta in your bloodstream than MG53, you scar easily.
Ma’s goal is to develop a therapy that will inhibit TGF Beta and promote MG53. Medical professionals can use the therapy during procedures to promote quick, scarless healing. His next step is to identify a small compound that can do this and eventually test whether it has the desired effect in human trials.
Presentation #2907, “MG53 promotes wound healing and reduces scar formation by facilitating cell membrane repair and controlling myofibroblast differentiation,” is authored by Haichang Li, Pu Duann, Pei-Hui Lin, Li Zhao, Zhaobo Fan, Tao Tan, Xinyu Zhou, Mingzhai Sun, Matthew Sermersheim, Hanley Ma, Steven Steinberg, Hua Zhu, Chunyu Zeng, Jianjun Guan and Jianjie Ma. It will be in a poster session that begins at 10:30 a.m. on Wed., March 2, 2016 in the West Hall of the Los Angeles Convention Center.

New method reveals high similarity between gorilla and human Y chromosome
Posted by Acubiz | BlogJim (on the right), whose Y chromosome was sequenced, together with Dolly, his mother, and Binti, his sister.
Credit: San Diego Zoo Global
A new, less expensive, and faster method now has been developed and used to determine the DNA sequence of the male-specific Y chromosome in the gorilla. The technique will allow better access to genetic information of the Y chromosome of any species and thus can be used to study male infertility disorders and male-specific mutations. It also can aid in conservation genetics efforts by helping to trace paternity and to track how males move within and between populations in endangered species, like gorillas.
A paper describing the method and the discovery resulting from its use in comparing the sequence of the gorilla Y chromosome to the sequences of the human and chimpanzee Y chromosomes will be published on March 2, 2016 in the Advance Online edition of the journal Genome Research. The article also will be published in the April 2016 print issue of the journal.
“Surprisingly, we found that in many ways the gorilla Y chromosome is more similar to the human Y chromosome than either is to the chimpanzee Y chromosome,” said Kateryna Makova, the Francis R. and Helen M. Pentz Professor of Science at Penn State and one of two corresponding authors of the paper. “In regions of the chromosome where we can align all three species, the sequence similarity fits with what we know about the evolutionary relationships among the species — humans are more closely related to chimpanzees. However, the chimpanzee Y chromosome appears to have undergone more changes in the number of genes and contains a different amount of repetitive elements compared to the human or gorilla. Moreover, a greater proportion of the gorilla Y sequences can be aligned to the human than to the chimpanzee Y chromosome.”
The Y chromosome of mammals is incredibly difficult to sequence for a number of reasons. One reason is that the Y chromosome is present in only one copy and makes up only about one to two percent of the total genetic material found in a cell of a male. To reduce this difficulty, the researchers used an experimental technique called flow-sorting to preferentially select the Y chromosome for sequencing based on the chromosome’s size and genetic content.
“Flow-sorting increased the amount of the Y chromosome in our dataset to about thirty percent,” said Paul Medvedev, assistant professor of computer science and engineering and of biochemistry and molecular biology at Penn State, the other corresponding author of the paper. “To further enrich our data for the Y chromosome, we developed a computational technique — called RecoverY — to sort the data into Y and non-Y sequences based on how frequently similar sequences appeared in our data.”
The Y chromosome, like all DNA, is composed of a series of molecules called “bases” that are represented by the letters A, T, C, and G. Current genetic sequencing technologies produce “reads” of sequence that are much shorter than the entire length of the chromosome. These reads need to be placed in order and pieced together by finding places where they overlap into longer and longer chunks. The research team used two different sequencing technologies to help with this assembly of the DNA sequence of the Y chromosome.
One sequencing technology used by the researchers produces massive amounts of very short reads — about 150 to 250 bases in length. Using this method, the researchers sequenced enough reads to cover the entire length of the Y chromosome about 450 times. The researchers assembled these short reads into longer chunks that they then further connected using the second sequencing technology that produces longer reads — about seven thousand bases in length on average.
“By reducing non-Y chromosome reads from our data with flow sorting and the RecoverY technique that we developed, and by using this combination of sequencing technologies, we were able to assemble the gorilla Y chromosome so that more than half of the sequence data was in chunks longer than about 100,000 bases in length,” said Medvedev.
Another reason that determining the genetic sequence of the Y chromosome is so difficult is that it is composed of an unusually high number of repeated sequences — regions where the sequence of As, Ts, Cs, and Gs are identical, or nearly identical, for thousands or millions of bases in a row. Many of these repeats, including some genes, appear as back-to-back series of the same repeated sequence or as long palindromes which, like the word “racecar,” read the same forward and backward. The researchers used an experimental technique — “droplet digital polymerase chain reaction” — to determine the number of copies of the genes that appear in these series.
“Sequencing the Y chromosome is like trying to put together a jigsaw puzzle, without knowing the final picture, from a pile of pieces where only about one out of every hundred is useful, and most of the pieces you do need look identical,” said Makova. “We’ve developed a pipeline for sequencing the Y chromosome that is more efficient than previous methods and reduces a number of the difficulties associated with determining the genetic sequence of the Y chromosome. Our method will open the door for studying the Y chromosome for more labs, more species, and more individuals within those species.”
To demonstrate the utility of the gorilla Y chromosome sequence they generated, the researchers designed genetic markers that can be used to differentiate the genetic relatedness among male gorillas and thus to aid in conservation genetics efforts targeted at preserving this endangered species.
In addition to Makova and Medvedev, the research team includes Marta Tomaszkiewicz, Samarth Rangavittal, Monika Cechova, Rebeca Campos-Sanchez, Howard W. Fescemyer, Robert Harris, Danling Ye, and Rayan Chikhi at Penn State; Malcom A. Ferguson-Smith and Patricia C. M. O’Brien at the University of Cambridge in the United Kingdom; and Oliver Ryder at the San Diego Zoo.
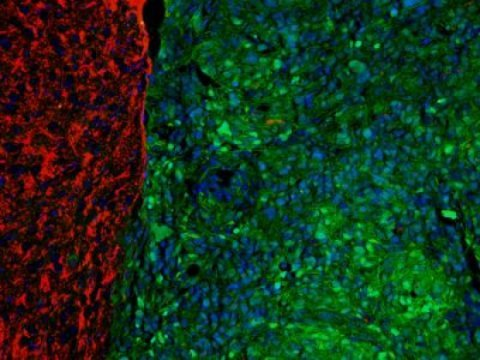
New weapon in the fight against children’s brain tumors
Posted by Acubiz | BlogTo create the new mouse model of children’s brain cancer, the researchers caused a mutation that occurs in many tumors, and used special stains to see the difference between healthy brain tissue (red) and the areas where the mutation took hold and a tumor arose (blue/green).
Credit: University of Michigan
Children with brain cancer may soon get some help from mice with the same disease, thanks to new research from University of Michigan Medical School scientists and their colleagues.
In a new paper in Science Translational Medicine, the U-M team describes how they developed a novel brain tumor model in mice.
The mice have the same genetic problems as those seen in many children with the most dangerous forms of brain cancer. That means the mice should be able to serve as a new test bed for treatments aimed at shrinking children’s tumors.
Unlike adults with brain cancer, children cannot receive radiation therapy, so doctors must rely on medications and other strategies. Brain tumors are the leading cause of cancer death in children. A lack of treatment options keeps survival rates low.
“This is exciting because it’s the first animal model of pediatric high-grade gliomas, or malignant brain tumors,” says Maria Castro, Ph.D., senior author of the paper and a professor in the departments of Neurosurgery and Cell and Developmental Biology at U-M. “The mice carry the genetic mutations found in human tumors, and develop tumors that closely resemble what children and adolescents do.”
And unlike previous attempts by others, the model generated at the U-M has a fully functional immune system, which makes it even more alike the children they mimic.
Brain cancer biology
In addition to developing the new genetically engineered mouse model for the form of cancer called glioblastoma multiforme, the researchers made a key discovery about brain tumor biology via the mice. Their work focuses on a protein called ATRX, and its role in helping cells repair damage to DNA. About one-third of children and young adults with brain cancer have ATRX mutations in their cancer cells.
Using a special genetic technique that introduced the same mutation in mice, soon after birth, the scientists were able to generate brain tumors that made less of the ATRX protein. Then, they showed that when the cancer cells didn’t make enough ATRX, the cells couldn’t join together the two ends of a broken DNA strand.
This genetic instability accelerated tumor growth and reduced the survival of mice that went without treatment.
But in an ironic — and potentially beneficial — twist, the researchers also showed that if they treated the mice’s tumors with drugs that damage DNA, they could actually shrink tumors more effectively and improve survival.
DNA-damaging cancer drugs already exist, and are used for other purposes in adults. If further research in the mice bears out, they could soon be tested in children whose brain tumors also show a decrease in ATRX production.
The first author of the study, U-M children’s cancer specialist Carl Koschmann, M.D., helps guide the treatment of children with brain cancer at U-M’s C.S. Mott Children’s Hospital. He also studies brain cancer at the basic cell level in the Castro laboratory.
“We are very excited about this tumor model as it mimics the developmental environment of a pediatric or adolescent human brain tumor,” he says. “We desperately need new therapies for pediatric GBM patients, as less than 20 percent of children diagnosed with GBM will survive five years. We currently base their treatment on a regimen designed for adult patients with GBM, which are very different tumors at the molecular level. Our mouse model has given us a great step towards developing targeted therapies to specific changes found in pediatric and adolescent GBM.”
Accelerating discovery
Dr Castro’s team at U-M, and their colleagues on teams at the Johns Hopkins School of Medicine, the University of Rochester, and the Institute of Cancer Research in London have been working toward this goal for several years.
A crucial aspect of the mouse model development was the ability to use what’s called the “Sleeping Beauty” technique of inserting genes into stem cells in the brain of newborn mice. This allows their brain to develop normally, but also triggers the development of cancer by adding mutated ATRX genes and other genes known to be involved in cancer.
They also looked at genetic data on several hundreds of adult and pediatric brain tumors from around the world. That helped them confirm that mutations in the gene that directs cells to make ATRX are responsible for making the tumors genetically unstable.
Personalizing treatment for brain tumors based on an individual’s ATRX mutation status is still a concept, not a reality. But the researchers believe their mouse model and the close collaboration with pediatric brain cancer treatment teams at Mott Hospital will accelerate the ability to do so.
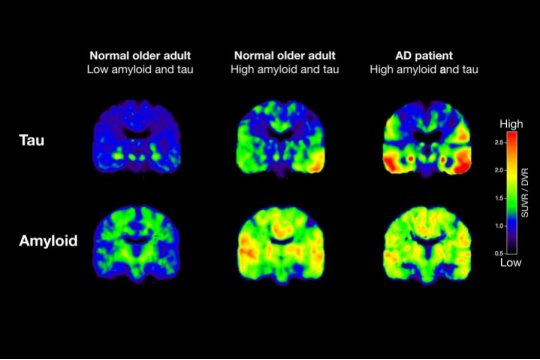
PET scans reveal key details of Alzheimer’s protein growth in aging brains
Posted by Acubiz | BlogShown are PET scans that track tau (top row) and beta-amyloid from two normal older people and a patient with Alzheimer’s disease (AD). The normal older adult on the left has no brain amyloid deposition and minimal tau in the medial temporal lobe. In the normal older adult in the middle, amyloid deposition is present throughout the brain, and tau has spread out into the temporal cortex. In the AD patient, both amyloid and tau are spread through the brain.
Credit: Michael Schöll
New research led by scientists at the University of California, Berkeley, shows for the first time that PET scans can track the progressive stages of Alzheimer’s disease in cognitively normal adults, a key advance in the early diagnosis and staging of the neurodegenerative disorder.
In the process, the scientists also obtained important clues about two Alzheimer’s-linked proteins — tau and beta-amyloid — and how they relate to each other.
The findings, to be published March 2 in the journal Neuron, come from positron emission tomography (PET) of 53 adults. Five were young adults aged 20-26, 33 were cognitively healthy adults aged 64-90, and 15 were patients aged 53-77 who had been diagnosed with probable Alzheimer’s dementia.
The stages of tau deposition were established by German researchers Heiko and Eva Braak through postmortem analysis of the brains of suspected Alzheimer’s patients.
“Braak staging was developed through data obtained from autopsies, but our study is the first to show the staging in people who are not only alive, but who have no signs of cognitive impairment,” said study principal investigator Dr. William Jagust, a professor at UC Berkeley’s School of Public Health and at the Helen Wills Neuroscience Institute and a faculty scientist at Lawrence Berkeley National Laboratory. “This opens the door to the use of PET scans as a diagnostic and staging tool.”
PET scans are used to detect early signs of disease by looking at cellular-level changes in organs and tissue. The results of the scans in this study paralleled Braak neuropathological stages, which range from 1 to 6, describing the degree of tau protein accumulation in the brain.
Jagust worked with study co-lead authors Michael Schöll, a visiting scholar, and Samuel Lockhart, a postdoctoral fellow, both at UC Berkeley’s Helen Wills Neuroscience Institute.
Tau vs. amyloid
Their findings also shed light on the nature of tau and amyloid protein deposits in the aging brain. For many years, the accumulation of beta amyloid plaques was considered the primary culprit in Alzheimer’s disease. Over the past decade, however, tau, a microtubule protein important in maintaining the structure of neurons, has emerged as a major player. When the tau protein gets tangled and twisted, its ability to support synaptic connections becomes impaired.
While a number of symptoms exist that signal Alzheimer’s disease, a definitive diagnosis has been possible only through an examination of the brain after the patient has died. The availability of amyloid imaging for the past decade has improved this situation, but how Alzheimer’s developed as a result of amyloid remains a mystery. Studies done in autopsies linked the development of symptoms to the deposition of the tau protein.
Through the PET scans, the researchers confirmed that with advancing age, tau protein accumulated in the medial temporal lobe — home to the hippocampus and the memory center of the brain.
“Tau is basically present in almost every aging brain,” said Schöll, who holds an appointment at Sweden’s University of Gothenburg. “Very few old people have no tau. In our case, it seems like the accumulation of tau in the medial temporal lobe was independent of amyloid and driven by age.”
The study revealed that higher levels of tau in the medial temporal lobe were associated with greater declines in episodic memory, the type of memory used to code new information. The researchers tested episodic memory by asking subjects to recall a list of words viewed 20 minutes earlier.
Both proteins involved in dementia
One question yet to be answered is why so many people who have tau in their medial temporal lobe never go on to develop Alzheimer’s. Likewise, adults may have beta amyloid in their brains and still be cognitively healthy.
“It’s not that one is more important than the other,” said Lockhart. “Our study suggests that they may work together in the progression of Alzheimer’s.”
While higher levels of tau in the medial temporal lobe were linked to more problems with episodic memory, it was when tau spread outside this region to other parts of the brain, such as the neocortex, that researchers saw more serious declines in global cognitive function. Significantly, they found that tau’s spread outside the medial temporal lobe was connected to the presence of amyloid plaques in the brain.
“Amyloid may somehow facilitate the spread of tau, or tau may initiate the deposition of amyloid. We don’t know. We can’t answer that at this point,” said Jagust. “All I can say is that when amyloid starts to show up, we start to see tau in other parts of the brain, and that is when real problems begin. We think that may be the beginning of symptomatic Alzheimer’s disease.”
What the study does indicate, the researchers said, is that tau imaging could become an important tool in helping to develop therapeutic approaches that target the correct protein — either amyloid or tau — depending on the disease stage.
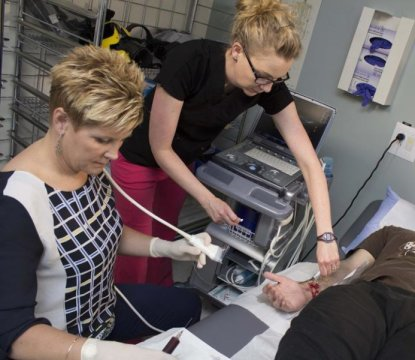
Platelet-rich plasma injections may lead to improvements in tissue healing
Posted by Acubiz | BlogDr. Marni Wesner performs a platelet-rich plasma injection procedure on a patient at the University of Alberta Glen Sather Sports Medicine Clinic.
Credit: University of Alberta
Tiger Woods, Kobe Bryant and A-Rod have all used it, but does platelet-rich plasma therapy (PRP) really work for the every-day active person? According to a University of Alberta Glen Sather Sports Medicine Clinic pilot study on patients with chronically sore shoulders published in PLOS ONE, preliminary findings say yes.
“We studied patients 35 to 60 years old with rotator cuff tendinopathy due to normal aging. For the first time, we were able to not only find reported improvements in pain and mobility, but also in the tissue — the MRI before and after showed structural change and a decrease in the size of tears,” says Marni Wesner, sports medicine physician at the Glen Sather Sports Medicine Clinic and lead author of the study.
Platelet-rich plasma injection (PRP) is an emerging therapeutic procedure in medicine and rehabilitation for the treatment of both acute and chronic soft tissue injuries. It involves collecting blood from the patient’s arm, separating the platelets via centrifuge and injecting it back into the patient’s injured tissue area to augment (or facilitate) the body’s natural healing response.
Platelets are cells that clot blood and contain over 300 active growth factors. In tissues that are aging and do not repair and regenerate well, growth factors may help to improve healing by creating an environment to foster healing and regeneration of tissue.
Though this is a pilot study on a small number of patients, the outcomes are still clinically relevant and this is the first time researchers have described structural change in the healing process as well as improvement in the patients’ pain and function.
“Based on MRI findings before and after the injections, we saw improvements in the tissue six months later in five of seven patients undergoing PRP and an appropriate rehabilitation program. The healing in the tissue appeared to correspond with the reported improvement of the pain and also with the clinical assessment of function,” explains Doug Gross, interim chair of physical therapy at the Faculty of Rehabilitation Medicine and corresponding author of the study.
For retired police officer Debbie Brown who is a participant in the study, this healing from PRP made all the difference in her life.
“For the past two years, I have tried everything for my right shoulder. Physio would help for a bit but then the problem would still be there. I tried acupuncture, Kinesio tape, cortisol injections — you name it, I’ve tried it,” says Brown. “Once I did the PRP, it really did fix everything!”
Brown says she did physical therapy with Heather Bredy at the Glen Sather Sports Medicine Clinic as part of her treatment and continues to do exercises regularly. Besides a bit of soreness here and there — she says she is 58, after all! — her shoulder is like new.
“I can shoulder-check now and brush my hair. I can work out and be active again,” she says.
Wesner says it’s important to note that to be considered for PRP at the Glen Sather Sports Medicine Clinic, the patient must be referred by a physician.
“The sports medicine physicians at the Glen Sather Sports Medicine Clinic always recommend and provide more conservative or ‘traditional’ care first, and appropriate rehabilitation along with the PRP injections” she says. “The Glen Sather Sports Medicine Clinic has done more than 600 PRP treatments on a variety of tendons and soft tissue areas. PRP injections at the clinic are ultra-sound guided.”
Gross, also the director of the Rehabilitation Research Centre, says PRP therapy is becoming more widespread and the next step in research is to study its effectiveness in larger, well-designed controlled clinical trials. “This pilot study has shown promising results and important experience for planning subsequent studies.”
As for Brown who’s had renewed function and healing in her shoulder, she can’t stop telling her friends and colleagues to go to the Glen Sather Clinic at the U of A.
“PRP makes me feel like I’m in my 20s!” she smiles.
Using graphene to fight bacteria
Posted by Acubiz | BlogNew research on graphene oxide may one day transform our ability to fight infections acquired in the hospital and elsewhere.
Scientists at the Università Cattolica del Sacro Cuore in Rome are studying graphene oxide in the hopes of one day creating bacteria-killing catheters and medical devices. Coating surgical tools with this carbon-based compound could kill bacteria, reducing the need for antibiotics, decreasing the rates of post-operative infections and speeding recovery times.
Graphene is a new wonder material first characterized in the last decade — work recognized with the 2010 Nobel Prize in physics. This single-atom-thin sheet material holds promise in future electronics, battery and manufacturing technologies.
“We want to make materials that will help patients and medical professionals,” said Valentina Palmieri a biotechnologist at the Università Cattolica del Sacro Cuore in Rome.
Graphene oxide, a form of graphene with molecular oxygen incorporated into it, protects against infection by destroying bacteria before it gets inside the body. The graphene oxide wraps around the bacteria, puncturing its membrane. A broken membrane prevents the bacteria from growing and often kills it. “The bacteria lose their complex structure and die,” Palmieri added. “And since graphene is just carbon — a building block of life — its cytotoxicity against human cells is much lower compared to any drug-based antimicrobial therapy.”
Researchers decided to use graphene oxide because it is very stable in a water solution, making it safe to interact with human cells. Graphene specifically attacks bacterial cells, while sparing human cells, and the mechanism behind this specificity is still unclear, Palmieri said. Current theories include that the material interacts more favorably with the bacterial cell wall or that mammalian cells have evolved multiple repair mechanisms to survive the chemical oxidation damage that graphene induces.
Graphene is also more eco-friendly. Traditional methods of preventing infection include antibiotic therapy and tools coated with silver — both of which are toxic to the environment, Palmieri said.
The compound is most effective when paired with salt. Too little salt and the graphene oxide won’t wrap around the bacteria. Too much salt and the graphene aggregates, failing to puncture the bacteria’s membrane. To destroy both Gram positive and Gram negative bacteria, two major categories of bacteria to which many hospital-acquired pathogens belong, a 300 nanometer sheet of graphene oxide solution must be mixed with low molarity (<10 mM) Calcium chloride.Palmieri and her team also created new protocol methods for testing the effectiveness of the graphene oxide and salt mixture. Researchers normally use a spectrophotometer to test whether the solution has killed all the bacteria. This method didn't account for any bacteria that may have fallen to the bottom of the solution. Instead, Palmieri uses two different wavelengths of light -- one that is sensitive to absorbing and one that is sensitive to scattering -- to check for bacteria.The next step is to test graphene oxide's effectiveness against fungi. Fungi can also cause significant problems if they infect an open wound. Up until this point, though, fungi cells are too big for graphene oxide to wrap all the way around them. Palmieri says she wants to alter the salt content inside the saline solution to see if she can solve this issue.
Energy drinks trigger abnormal heart rhythm, rise in blood pressure
Posted by Acubiz | BlogA clinical trial led by researchers from University of the Pacific and David Grant Medical Center adds to the evidence that energy drinks may be bad for your heart. Results of the study will be presented today at a meeting of the American Heart Association in Phoenix, Arizona.
“Our findings suggest certain energy drinks may increase the risk of having an abnormal heart rhythm when consumed in high volumes,” said primary investigator Sachin A Shah, an associate professor of pharmacy practice at Pacific’s Thomas J. Long School of Pharmacy and Health Sciences. “While we wait for more data, some consumers should exercise caution and not blindly follow the buzz.”
Phillip Oppenheimer, dean and professor of pharmacy practice at Pacific, said the findings are of special concern among young adults. “Energy drinks are widely consumed within the college population, which further extends the relevance of this study,” Oppenheimer said.
The study enrolled 27 healthy volunteers between the ages of 18 and 40. Subjects drank either two cans of an energy drink, an equivalent volume of a drink containing panax ginseng (an ingredient in the energy drink), or a placebo beverage once a day, every six days, for three weeks. Neither the volunteers nor the researchers knew who was getting which drink until the end of the three weeks.
The researchers measured subjects’ heart rhythm and blood pressure before the drinks were consumed and four times during the six hours immediately afterward.
The volunteers who consumed the energy drink experienced a statistically significant increase in a marker of abnormal heart rhythm risk known as the QTc interval. These volunteers also experienced a slight rise in blood pressure. These effects persisted for two hours after the energy drink was consumed.
In contrast, the ginseng and placebo groups showed no rises in QTc interval or blood pressure.
Shah noted that some drugs contain a warning in their package insert when the drug has been shown to prolong the QTc interval to a degree similar to that seen in the study (6 milliseconds).
Energy drinks have been associated with sudden deaths. As of June 2014, the Center for Science in Public Interest, a consumer health advocacy group, had collected reports of 34 deaths that may have been associated with energy drinks.
According to Shah, more research needs to be performed assessing the heart rhythm and blood pressure effects of energy drinks, especially in those with underlying cardiac conditions such as congenital long QT syndrome and hypertension.